The Stability Paradox – How Do Organisms Change Shape Over the Course of Evolution?
Researchers at the Technion have discovered how changes in genetic regulatory sequences can lead to alterations in the form and structure of animals – even when genetic regulatory systems are stable and resistant to change. The study, published in Science Advances, was led by Dr. Ella Preger-Ben Noon and Ph.D. candidate Areej Said-Ahmad from the Ruth and Bruce Rappaport Faculty of Medicine.
The loss of morphological traits is a common phenomenon in evolution. Well-known examples include the loss of legs in snakes and the loss of eyes in cavefish. In many cases, such changes do not result from the loss of the genes responsible for these traits, but rather from changes in how those genes are regulated during development. However, many developmental genes are controlled by multiple regulatory sequences with overlapping activity, forming a stable and robust regulatory system.
This study addresses a fundamental question in biology: how do organisms change form over the course of evolution despite the presence of stable genetic regulatory systems? These systems rely on DNA sequences known as enhancers, which activate genes at precise times, levels, and locations during development. Enhancers often act redundantly, so that if one is impaired, others can compensate and maintain proper gene expression. This redundancy confers stability and resistance to change, but also raises a paradox: how do changes in gene expression still occur, leading to alterations in the shape and structure of organs?
To address this question, the researchers focused on Drosophila flies, particularly the species Drosophila sechellia, in which tiny hair-like structures (trichomes) have disappeared from the larval body during evolution. This trait is controlled by the shavenbaby gene, whose expression is regulated by multiple enhancers. Contrary to expectations that such a system would protect gene expression from change, the researchers found that four different enhancers of shavenbaby lost their activity over the course of evolution, each through a distinct mechanism.
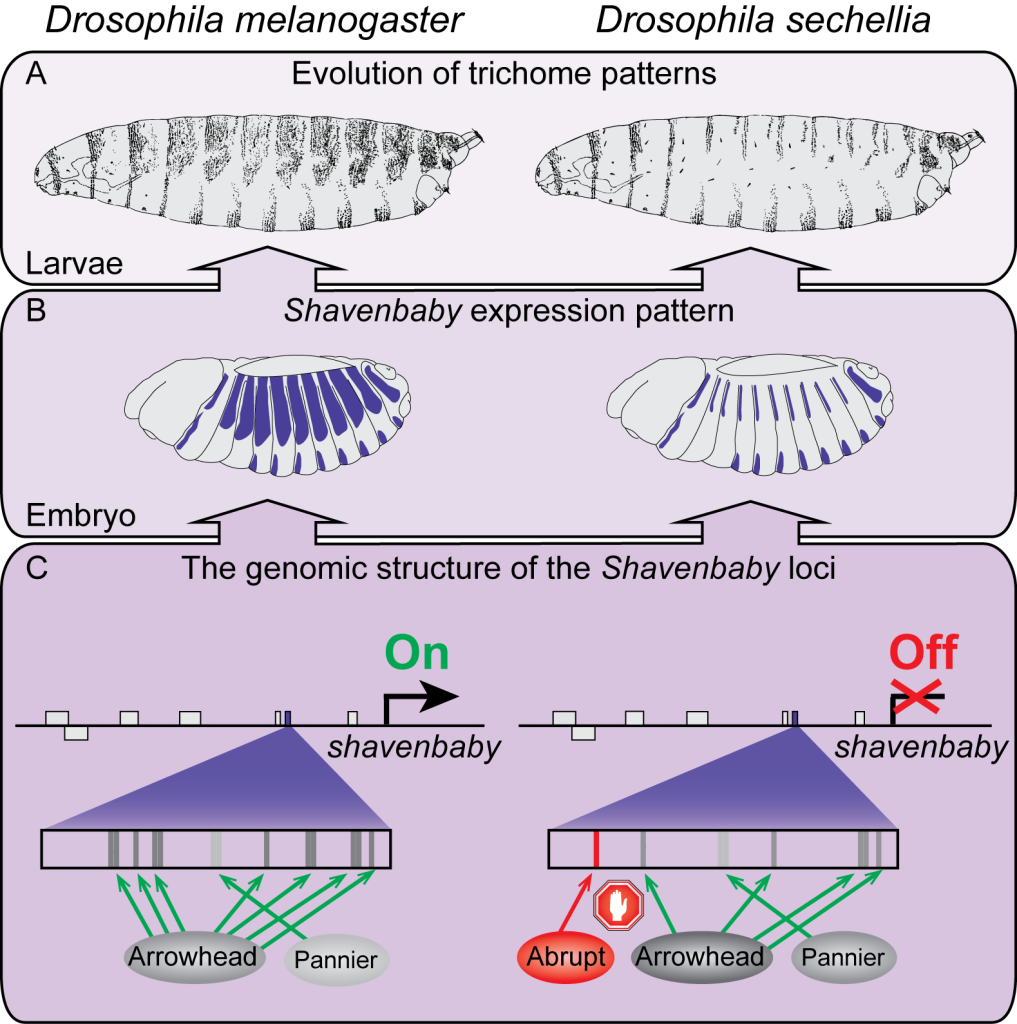
Through detailed DNA sequence analysis and functional experiments, the researchers found that the loss of enhancer activity occurred via different molecular mechanisms, including deletion of essential sequences, loss of binding sites for activators and gain of repressor binding sites, acquisition of a silencer, and even the unmasking of pre-existing repression. In other words, the same evolutionary outcome – the loss of gene expression – was achieved through different molecular pathways within the same genomic region.
These findings demonstrate that the same evolutionary outcome can arise through multiple routes. The presence of multiple enhancers, while they contribute to stable gene expression, also creates points of vulnerability where mutations can reduce their activity. The study shows that stability does not necessarily act as a barrier to evolution, as there are diverse molecular ways to circumvent it. These insights are relevant to a wide range of biological systems and deepen our understanding of how variation in form and structure arises in nature.
To read the full article, click here
Photo Credit: Rami Shelush