Ultralong DNA Through a Nanopore
Threading ultralong DNA through a nanopore
Researchers at the Technion–Israel Institute of Technology disentangled and threaded a DNA molecule that is hundreds of thousands of nucleobase base pairs long through a nanopore. This breakthrough is expected to facilitate the study of genomic DNA.

l-r: Dr. Diana Huttner, PhD student Adam Zrehen, Professor Amit Meller.
The nanopore device was developed by Professor Amit Meller, Dr. Diana Huttner, and PhD student Adam Zrehen of the Technion Faculty of Biomedical Engineering. Their pioneering research was described in a paper published in the November 2019 issue of ACS Nano.
Nanopore sensors are used to read the genetic code in DNA, which is composed of bases or “letters.” The sensor consists of a nanoscale pore about 1/10,000 the diameter of human hair, through which a single DNA is threaded and read. While reading short DNA strands such as that of viruses is routine, it is especially difficult to thread intact ultralong human DNA through such a tiny aperture because DNA forms a random coil in solution. The process can be likened to threading tangled yarn through a needle.
Prof. Meller and his team successfully disentangled and threaded a DNA molecule that is hundreds of thousands of base pairs long through a nanopore. The DNA was labelled by using light-emitting dyes so that it can be tracked and manipulated in real-time by pressure and electric fields in a glass-sealed silicon chip about the size of a coin.
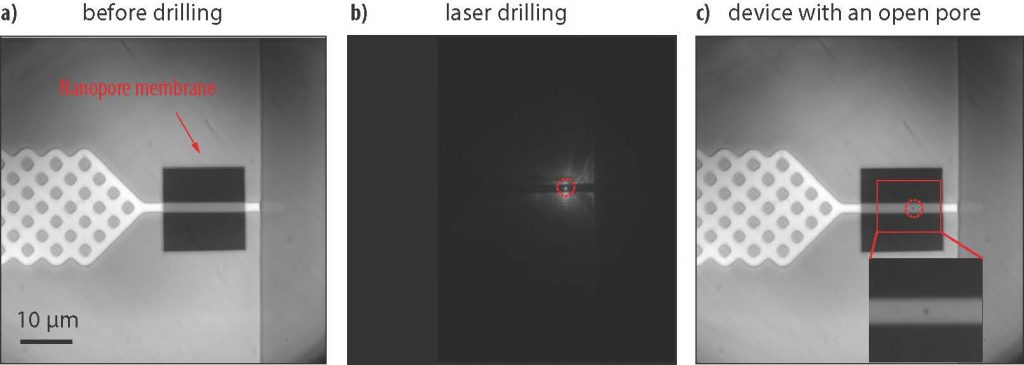
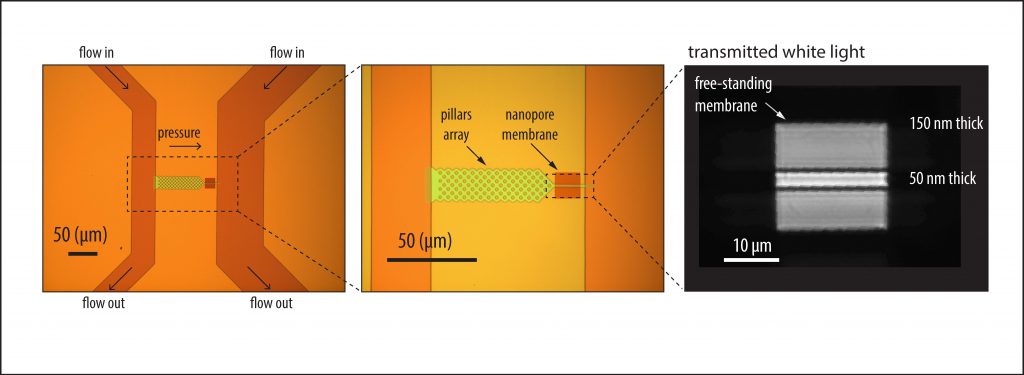
The device developed by the researchers is designed for simultaneous electrical and optical detection and sorting of ultralong (~500 kilobase pair) genomic DNA. Fabricated in silicon, it features a central pillar array for stretching the ultralong DNA and a narrow channel for funneling the linearized molecule to the nanopore. The nanopore is “drilled” directly in the channel by using a focused laser that controllably etches minute amounts of material. The low height profile of the device permits high numerical aperture and high magnification imaging of individual DNA molecules.
As the first direct visualization of DNA threading through a nanopore while recording its electrical signal, the study furthers the understanding of the DNA capture and threading process,. It also promotes the development of all-in-one micro/nanofluidic platforms for nanopore sensing of biomolecules. Extremely low concentrations (50 fM) and sample volumes (∼1 μL) of DNA can be processed, making the device highly compatible with clinical samples.
Since reading long, intact DNA is less prone to error compared with assembling many small DNA fragments, the researchers anticipate their design will play an important role in studying genomic DNA.
The project received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program and was also supported by the i-Core program of the Israel Science Foundation.
Click here for the paper in ACS Nano.